Chapter 3 Developmental biology of the eye
Introduction
For both ethical and technical reasons, experimental developmental biology is not possible in humans. This limitation is likely to change with the ability to induce trans-differentiation of adult human cells into induced pluripotent stems (iPS) cells.1 An ever-increasing understanding of the genetic basis of human malformations combined with the availability of patient-derived iPS cells predicts that human developmental biology will be an important and rapidly growing field. However, our current knowledge of eye development comes from Drosophila and vertebrate models: frogs (Xenopus laevus, Xenopus tropicalis), fish (zebrafish: Danio rerio, medaka: Oryzias latipes), chick (Gallus gallus), and mouse (Mus musculus). Each has strengths and weaknesses. For example, chick embryos have been extensively used for fate mapping (Box 3.1) and tissue recombination experiments, but there are few natural mutations available for study and genetic manipulation is difficult. In mice, it is possible to inactivate almost any gene in a targeted manner via homologous recombination (see Box 3.1) in embryonic stem cells, but very early development is difficult to visualize since this is a placental mammal. Although exact equivalence of animal model experimental data with orthologous processes in humans is unlikely given the expansion of gene families through evolutionary genome duplication and existence of species-specific phenomena, it is likely that many of the developmental mechanisms will be common and generalizable.
Box 3.1
Definition of terms
Domain − a specific region or amino acid sequence in a protein with a particular function
Fate mapping − a technique developed by Vogt to trace the specific regions of an early embryo
Ligand − a signal-triggering molecule that binds to a site on a target protein
Paralogue − a pair of genes that derive from the same ancestral gene
Important concepts and processes in developmental biology
Cell migration
Embryogenesis involves dramatic shape changes over short periods of time. Much of this change is due to differential growth rates within and between tissues. However, there is a subset of cells that shows remarkable migration from their birth place to distant regions of the embryos. The best known of these traveling cells are neural crest cells which form on the lips of the neural fold along the length of the neural tube. The cranial neural crest cells are important in eye development. The molecular basis of the movement of neural crest cells is evolving and many of the master control genes for this process are also important in mediating cancer invasion and metastasis.2
Programmed cell death
In many developmental tissues a proportion of the constituent cells will be seen to undergo a process where there is nuclear fragmentation leading to death. This process is microscopically and molecularly distinct from senescence and is known as apoptosis or programmed cell death (PCD).3 The correct functioning of PCD is crucial to the normal development of many organs and appears to prune excess cells.
Signaling
Signaling is one of the most important processes in developmental biology. It involves interaction of a ligand (see Box 3.1) with a receptor, which then effects a change, usually altering protein phosphorylation and/or the transcriptional (see Box 3.1) profile of the receptor-bearing cell. Ligands can be proteins (e.g. Sonic hedgehog) or small molecules (e.g. all-trans-retinoic acid). The receptors (e.g. fibroblast growth factor receptors) can be on the cell surface or intracellular (e.g. retinoic acid receptors). The ligand−receptor interaction often leads to a signal transduction (see Box 3.1) cascade (e.g. MAPK pathway). Some ligand−receptor interactions activate several different signal transduction pathways when the most commonly used pathway is known as canonical (e.g. beta catenin activation in Wnt signaling) and the alternative pathways known as non-canonical. There are many different signaling pathways used during eye development (Fig. 3.1).
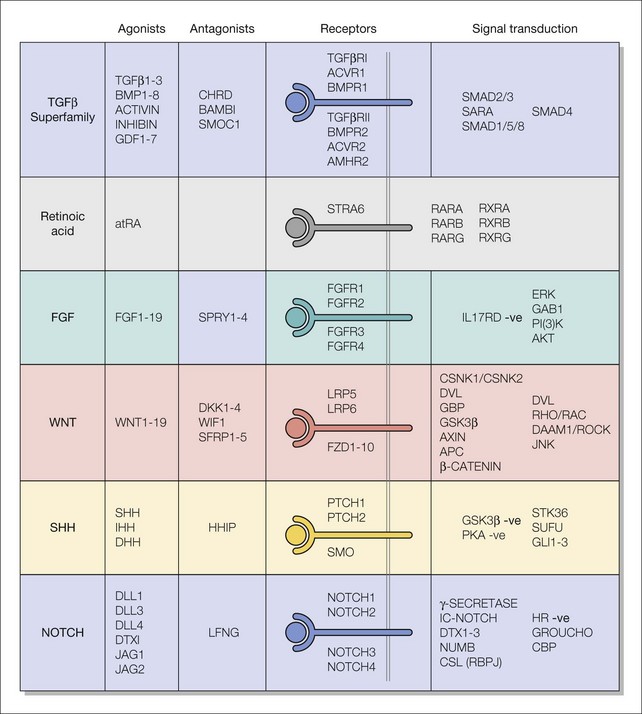
Fig. 3.1 Major signaling pathways used during eye development.
The signaling pathways are made up of ligands, antagonists, receptors, and signal transduction effectors. The pathways are named after the ligands, which are either groups of proteins that are related by sequence homology (see Box 3.1) or small molecules. Some receptor−ligand interactions may use more than one signal transduction cascade depending on cell context.
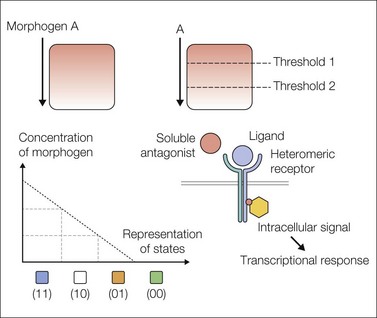
Morphogen gradients (see Box 3.1) involve a diffusible ligand exerting concentration-dependent differential effects on target tissues. This important concept is summarized in the so-called French flag model (Fig. 3.2).